The following list includes a few examples of BIDAC expertise areas and software tools. The list is neither complete nor exclusive, and researchers are advised to consult the website of the SCI Institute (www.sci.utah.edu) for additional information on research projects, publications and software tools and methods.
- AI-based analysis
- Animal image analysis
- Microscopy analysis
- NeuroImage analysis
- Segmentation analysis
- Software development
- General Infrastructure and computational resources
AI-based analysis
We have developed expertise in building, training and comparing state-of-the-art Convolutional Neural Networks (CNN) and Deep Neural Networks (DNN) to enable robust image classification, regression and segmentation tasks.
Examples of project collaborations entail:
- Deep learning analysis for image classification and regression tasks
- 3D ranging from thermal camera systems via deep neural networks
- Brain aneurysm identification and segmentation on 3D TOF-MRA acquisitions via neural networks
Animal Image Analysis
Mario Capecchi (CIBC)
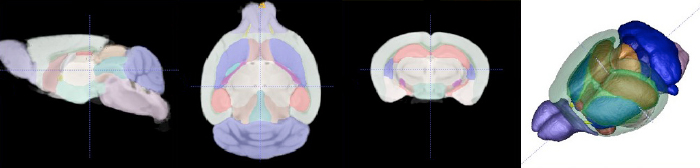
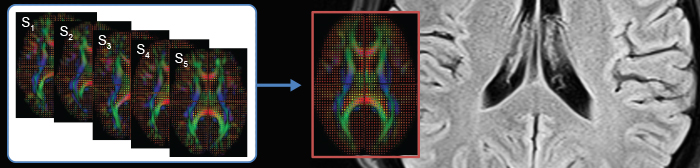
Images of the Utah mouse imaging dataset (images courtesy Naveen Nagarajan, Capecchi Lab) illustrating automatic segmentation of brain tissue and anatomical structures via the 3D Paxinos atlas and brain connectivity analysis of diffusion-weighted MRI (DW-MRI). The BIDAC provided expertise in MRI pulse sequences for in-vivo and ex-vivo mouse brain studies optimized for automated quantitative image processing and statistical evaluation.
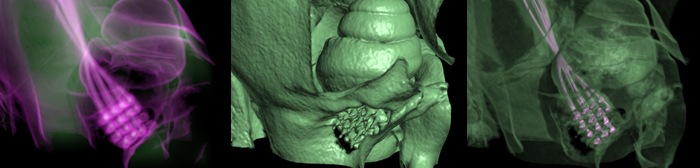
Richard Normann (Imaging of Utah Electrode Array, Implanted in Cochlear Nerve)
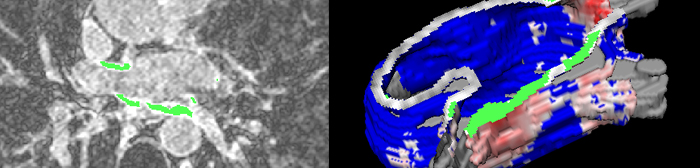
In spite of its outstanding success in some profoundly deaf individuals, cochlear implants are not effective in all implanted subjects. Problems of limited low frequency restoration, limited numbers of channels, and high stimulation threshold currents are a result of the stimulating electrodes being located some distance away from the cochlear nerve, with the modiolar bone intervening between the electrodes and the nerve. An array of penetrating electrodes, inserted directly into the auditory nerve may mitigate some of these problems. The active tips of the implanted electrodes abutting the cochlear nerve fibers should allow much more focal stimulation than can be achieved via present cochlear electrode arrays. The Utah Electrode Array has been surgically implanted into the cochlear nerve of felines for periods of over six months. Non-invasively verifying the accuracy of the electrode array's placement within the cochlear nerve, in an anatomical region completely encased in thick temporal bone, requires a combination of high-resolution scanning, volumetric image processing, and visualization techniques.
Microscopy Analysis
Fluorender
FluoRender is an interactive rendering tool for confocal microscopy data visualization. It combines the renderings of multichannel volume data and polygon mesh data; the properties of each dataset can be adjusted independently and quickly. The tool is designed especially for neurobiologists, and helps them better visualize the fluorescent-stained confocal samples. The FluoRender development package includes tools for 3D confocal microscopy file conversion. Find out more on the FluoRender's website.
Robert Marc (Connectomics and Automatic Mosaicking)
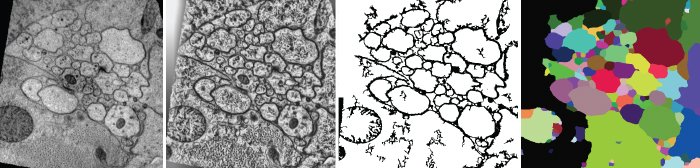
Detection of neuron membranes in electron microscopy images
Studying nervous systems via the connectome, the map of connectivities of all neurons in that system, is a challenging problem in neuroscience. In working towards a solution, neurobiologists have acquired large-electron microscopy datasets. However, the sheer volume of these datasets renders manual analysis nearly impossible. Hence, automated image analysis methods are required for reconstructing the connectome from these very large image collections. Segmentation of neurons in these images, an essential step of the reconstruction pipeline, is also difficult because of noise, anisotropic shapes and brightness, and the presence of confounding structures. The method we've developed uses a series of artificial neural networks (ANNs) in a framework combined with a feature vector that is composed of image intensities sampled over a stencil neighborhood. Several ANNs are applied, allowing each ANN to use the classification context provided by the previous network to improve detection accuracy. We've developed the method of serial ANNs to show that the learned context improves detection over traditional ANNs. We also present specific advantages over previous membrane-detection methods. The results are a significant step towards an automated system for the reconstruction of the connectome.
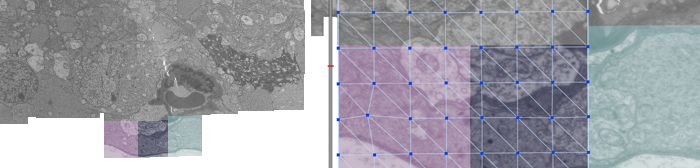
Automatic mosaicking and volume assembly of Microscopy Images
We present a computationally efficient, robust, and fully automatic method for large-scale electron and confocal microscopy image registration. The proposed method is able to construct large image mosaics from thousands of smaller, overlapping tiles with unknown or uncertain positions, and then align sections from a serial section capture into a common coordinate system. The method also accounts for nonlinear deformations, both in constructing sections and in aligning sections to each other. The publicly available software tools include the algorithms and a Graphical User Interface for easy access to the algorithms.
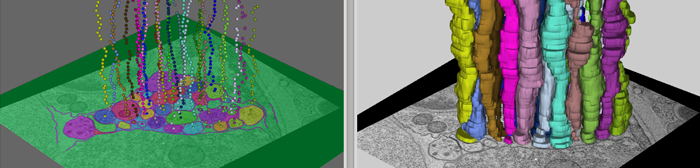
Nerve (Semi-automated Neuron Boundary Detection)
Neuroscientists are developing new imaging techniques and generating large volumes of data in an effort to understand the complex structure of the nervous system. The complexity and size of this data makes human interpretation a labor-intensive task. To aid in the analysis, new segmentation techniques for identifying neurons in these feature rich datasets are required. We have developed a method for neuron boundary detection, segmentation, and visualization. It combines both automated segmentation techniques with a graphical user interface for correction of mistakes in the automated process. This combination enables users to quickly segment cellular processes in large volumes.
Neuroimage Analysis
Brain Atlas
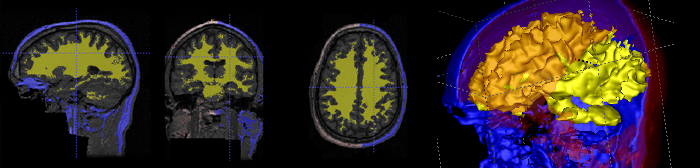
Construction of population atlases is a key issue in medical image analysis, and particularly in brain mapping. Large sets of images are mapped into a common coordinate system to study intra-population variability and inter-population differences, to provide voxel-wise mapping of functional sites, and help tissue and object segmentation via registration of anatomical labels. A Driving application has been primarily child neuroimaging autism, and recent research focuses on large sets of brain images.
Research Projects
Alzheimer's Disease Neuroimaging Initiative
Brain Population Analysis with Manifold Models
Non-parametric Brain Atlas Building
Segmentation and Analysis
Corview
Corview is a clinical image-analysis software developed at CARMA that implements late gadolinium enhancement MRI post processing workflows for AF management. Corview incorporates innovate image-processing algorithms and MRI volume visualization to produce detailed patient-specific models of heart structures, assess left-atrial tissue structural remodeling for AF staging, and assess the regional patterning and extent of post-radiofrequency ablation scarring. In support of other CARMA research efforts, Corview also serves as a platform for rapid prototyping and deployment of new cardiac image analysis. At CARMA, the unique embedding of dedicated software architects and image-processing scientists in the hospital environment has produced a streamlined tool for clinical work that has been designed, in part, by the clinicians themselves. Corview is currently in early release within the CARMA center and in daily use by CARMA MRI processing technicians.
ITK-SNAP
ITK-SNAP: Is a free open-source multiplatform (Linux, Mac OS, Windows) software application based on the Insight Toolkit computational environment used to segment structures in 3D (bio)medical images. It provides semiautomatic segmentation using active contour methods, manual delineation and image navigation and a large set of supporting utilities.
Software development
Specific driving biomedical and biological applications require adaptation of tools and methods to specific local needs, but in turn lead to updates of the respective software packages available to the whole research community. This activity can include data format conversions, adaptation to new/different operating systems and integration of software packages into specific computational platforms.
BIDAC staff and associated researchers have strong expertise in all aspects of image reformatting, image preprocessing, filtering, image registration (same or multiple modalities), interpolation and image segmentation based on statistical pattern recognition, active contours and machine learning concepts. More information on the software can be found in the Software page, or on the SCI Institute Software page.
General infrastructure and computational resources
BIDAC is housed at the University of Utah's Scientific Computing and Imaging Institute. BIDAC staff and researchers have access to excellent computational infrastructure, 500+ Terabytes of data storage and high-bandwidth connectivity. We have installed common computational tools used by the scientific community, including Matlab, Mathematica, Insight Toolkit and tools dedicated to specific image processing and visualization tasks. Meetings with BIDAC staff members can be arranged at the SCI Institute, including short-term guest visits for training on use of software applied to the researchers' own datasets.